GcDUO: A Batch Processing Framework Integrating PARAFAC and PARAFAC2 for GC×GC-MS Data Analysis (Maria Llambrich, MDCW 2026)

- Photo: MDCW: GcDUO: A Batch Processing Framework Integrating PARAFAC and PARAFAC2 for GC×GC-MS Data Analysis (Maria Llambrich, MDCW 2026)
- Video: LabRulez: Maria Llambrich: GcDUO: automating GC×GC-MS data analysis via PARAFAC and PARAFAC2 (MDCW 2026)
🎤 Presenter: Maria Llambrich (Institut d’Investigació Sanitària Pere Virgili)
Abstract
Multidimensional chromatography-mass spectrometry (MS) is a powerful analytical technique that integrates two or more chromatographic separations with MS, offering superior resolution, increased signal-to-noise, and selectivity for complex sample analysis. Despite its potential, its adoption remains limited due to data complexity and processing challenges. Chemometric approaches, particularly multiway models like Parallel Factor Analysis (PARAFAC), have proven effective in addressing these challenges by enabling the extraction of meaningful chemical information from multidimensional datasets. However, traditional PARAFAC is constrained by its assumption of data tri-linearity, which may not be valid in all cases, where data have misalignments. To overcome these limitations, we present GcDUO, an open-source data processing software that enables annotation, deconvolution, and analysis of batch GC×GC-MS data (Llambrich et al., Briefings in Bioinformatics 2025, doi: 10.1093/bib/bbaf080, https://github.com/mariallr/GcDuo).
GcDUO, implemented in R, accepts non-vendor-specific standardized CDF files, and rearranges the data into four-dimensional tensor structures, preserving the GC×GC-MS data structure. GcDUO integrates advanced chemometric methods, including PARAFAC and PARAFAC2, for a more accurate and comprehensive analysis. PARAFAC is particularly useful for deconvoluting overlapping peaks and extracting pure chemical signals, while PARAFAC2 relaxes the tri-linearity constraint, allowing batch analysis for samples. GcDUO achieves both high-resolution peak detection and robust quantification across complex GC×GC-MS datasets. The software was validated against the gold-standard software for comprehensive GC, demonstrating a high correlation (R² = 0.9) in peak area measurements, confirming its effectiveness and reliability. GcDUO provides a valuable, open-source platform in the comprehensive chromatography field, enabling more accessible and customizable data analysis.
Video Transcription
Gas chromatography coupled with comprehensive two-dimensional separation (GC×GC), especially when combined with high-resolution mass spectrometry (HRMS), has become a widely used technique for the analysis of highly complex samples. It provides enhanced chromatographic resolution, increased peak capacity, and improved sensitivity, enabling more reliable detection and compound annotation.
These advantages have driven adoption across multiple application areas, including:
- forensic chemistry
- environmental analysis
- petrochemical and fuel characterization
- food, flavors, and fragrances
- biological applications such as metabolomics and breath analysis
Despite these benefits, increasing instrumental performance also leads to significantly higher data complexity.
Data Complexity in GC×GC–MS
A single GC×GC–MS experiment produces third-order data consisting of:
- first-dimension retention time
- second-dimension retention time
- mass spectral data
When multiple samples are analyzed, the dataset becomes four-dimensional, forming a complex tensor structure.
While this enables advanced chemometric analysis, it also creates a barrier for users, as:
- data processing requires specialized workflows
- interpretation demands chemometric expertise
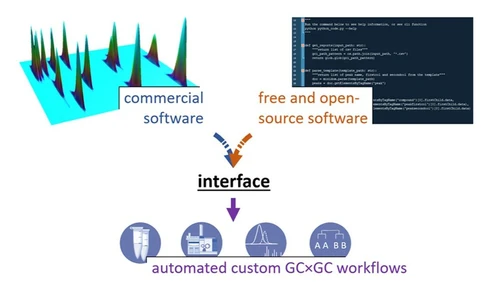
- commercial software often lacks flexibility and automation
GC Duo: Open-Source Data Processing Tool
To address these challenges, an open-source software tool called GC Duo was developed.
Key characteristics
- Implemented in R
- Available on GitHub
- Designed for untargeted and batch analysis
- Processes raw GC×GC–MS data without prior knowledge of samples
Core functionalities
- feature detection
- deconvolution
- spectral identification
- batch processing
Data Processing Workflow
1. Data Import and Structuring
- Reads standard CDF files (vendor-independent format)
- Converts raw data into 4D tensor structures using modulation parameters
2. Region of Interest (ROI) Selection
- Uses an inverse watershed algorithm
- Identifies significant signal regions based on:
- peak shape
- signal-to-noise ratio
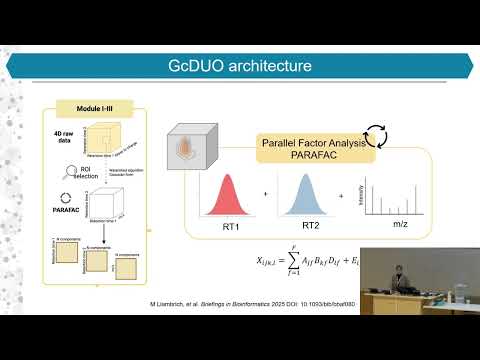
3. Feature Extraction via PARAFAC
- Applies Parallel Factor Analysis (PARAFAC)
- Decomposes data into:
- chromatographic profiles (1st and 2nd dimension)
- corresponding mass spectra
- Iterative modeling determines the optimal number of components
4. Feature Consolidation
- Removes duplicates across samples
- Generates consensus spectra
- Matches spectra against libraries using similarity scoring
- Optional use of retention indices
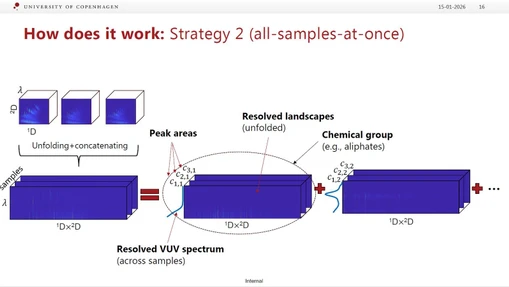
5. Refinement Step (Key Innovation)
- Re-applies the model using:
- all samples simultaneously
- defined retention windows
- predefined number of components
- Uses PARAFAC2 for batch data to handle deviations from trilinearity
Outcome
- improved spectral quality
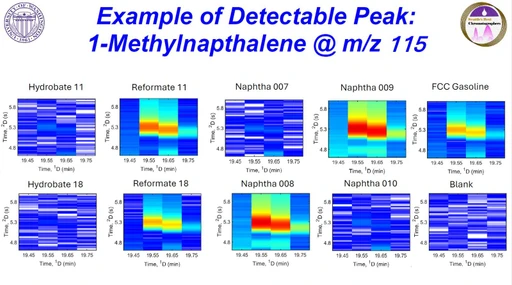
- detection of low-intensity or unresolved peaks
- more accurate peak area calculation
Results and Validation
Training Dataset
- Mixture of 36 compounds at varying concentrations
- ROI detection successfully identified relevant chromatographic regions
- PARAFAC improved spectral clarity and library matching
PARAFAC vs. PARAFAC2
- PARAFAC may distort peak alignment → potential quantification errors
- PARAFAC2 preserves original peak shapes → more accurate peak areas
Validation Datasets
1. Fruity Beer Dataset
- 27 compounds
- 85% successfully annotated
- Missing detections due to:
- absence in library
- low signal intensity
2. Breath Mix Dataset
- 12 compounds
- 100% identification achieved (with adjusted S/N threshold)
- Lower thresholds increased noise → trade-off observed
Performance Comparison
- Compared with vendor software (ChromaTOF)
- Results:
- consistent retention times
- strong correlation of peak areas (r ≈ 0.9)
Additional Observations
- inherent noise reduction through PARAFAC
- elimination of additional preprocessing steps
Conclusions
GC Duo provides a flexible and comprehensive solution for GC×GC–MS data processing.
Key advantages
- open-source and customizable
- vendor-independent data handling
- advanced chemometric tools (PARAFAC, PARAFAC2)
- improved reproducibility and interpretability
Limitations
- high computational demand
- sensitivity to data linearity assumptions
- requires initial data quality assessment
Final Remarks
GC Duo represents a powerful platform for untargeted GC×GC–MS studies, enabling advanced data analysis while maintaining flexibility and transparency. Its open-source nature encourages community-driven development and optimization.
-Workshop-LOGO_s.webp)