Tile-based Analysis of GC×GC-TOFMS Data of SPME Sampled VOCs Produced from Pseudomonas aeruginosa and Aspergillus fumigatus (Wenjing Ma, MDCW 2026)

- Photo: MDCW: Tile-based Analysis of GC×GC-TOFMS Data of SPME Sampled VOCs Produced from Pseudomonas aeruginosa and Aspergillus fumigatus (Wenjing Ma, MDCW 2026)
- Video: LabRulez: Wenjing Ma: Tile-based Fisher-ratio analysis of GC×GC-TOFMS data of SPME sampled VOC (MDCW 2026)
🎤 Presenter: Wenjing Ma (University of Washington)
Abstract
Pseudomonas aeruginosa and Aspergillus fumigatus are major pathogens found in the lungs of Cystic fibrosis patients. Their coexistence worsens lung function and leads to poor clinical outcomes. To investigate their metabolic interactions, we analyzed the volatile space of P. aeruginosa and A. fumigatus using SPME-GC×GC-TOFMS across four sample classes: Media, P. aeruginosa monoculture, A. fumigatus monoculture, and their co-culture. GC×GC-TOFMS provides high-resolution, high-sensitivity separation of complex metabolomic samples, but it also generates high-dimensional data that can be challenging to analyze.
Therefore, feature selection and chemometric methods are essential to extract meaningful information. Additionally, peak table alignment issues due to retention time shifting across samples can in principle hinder the comparative analysis of multiple samples. Hence, we applied tile-based Fisher-ratio analysis using ChromaTOF Tile software to discover analytes that are statistically significant in concentration differences across samples with replicates while minimizing retention-time shifts. This platform generates a comprehensive hit list that links detected analytes across all samples. In summary, our study integrated tile-based analysis to generate a comprehensive peak table that relates all analytes in all samples in terms of up and down regulation of their concentrations. With further analyte identification, we have characterized analytes specific to each sample class to ultimately learn more about Pseudomonas aeruginosa and Aspergillus fumigatus pathogens and their co-culture.
Video Transcription
In this study, Wenjing Ma from the University of Washington presents a data-driven approach for analyzing complex GC×GC–MS datasets, with a focus on volatile metabolites produced by clinically relevant microorganisms. The research centers on the interaction between Pseudomonas aeruginosa and Aspergillus fumigatus, two pathogens commonly found in the lungs of cystic fibrosis patients. Their co-existence is known to correlate with worse clinical outcomes, making the understanding of their metabolic interplay particularly important.
The main objective of the work is to identify metabolites that differ significantly between monocultures and co-cultures, thereby revealing biochemical interactions that may influence disease progression.
Experimental Approach
The experimental design includes four distinct sample classes: a media control, monocultures of Pseudomonas aeruginosa and Aspergillus fumigatus, and their co-culture. Samples were prepared using a microfluidic platform and analyzed via headspace sampling, enabling the capture of volatile compounds released during growth.
To characterize these samples, comprehensive two-dimensional gas chromatography coupled with time-of-flight mass spectrometry (GC×GC–TOF-MS) was employed. This technique provides high acquisition speed and full mass spectral information, allowing detailed chemical characterization. However, it also generates extremely complex datasets. A single chromatogram may contain up to a million data points, and when multiple samples are considered, the data effectively become four-dimensional. Such complexity makes direct manual interpretation impractical.
Data Processing Strategy
Traditional approaches to data reduction, such as generating peak tables or averaging chromatograms, present significant limitations. Peak alignment can be compromised by retention time shifts, while averaging can reduce resolution through peak broadening. To overcome these issues, the study applies a tile-based feature selection strategy built on statistical analysis.
The chromatographic space is divided into small regions, referred to as tiles, and each tile is evaluated using a statistical metric that compares variability between sample classes to variability within classes. This approach allows the identification of features that show the greatest differences across experimental conditions. Importantly, the method is extended to two-dimensional chromatographic data and applied across individual mass channels, enabling a highly detailed and systematic analysis.
The workflow is implemented using Chromatof Tile software, which produces ranked lists of statistically significant features and supports visualization tools such as principal component analysis (PCA) plots for confirming class separation.
Results and Interpretation
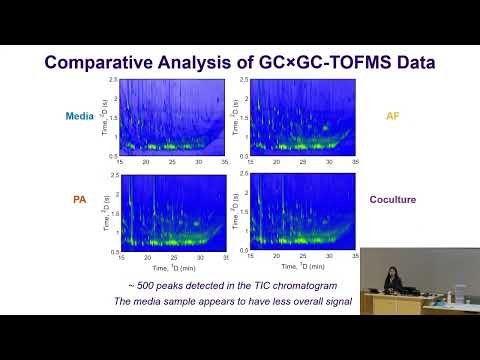
Initial inspection of raw chromatograms reveals that while the media control exhibits relatively low signal intensity, the biological samples show highly complex and overlapping patterns. Differences between monocultures and co-cultures are not readily apparent without advanced data processing.
Using tile-based analysis, hundreds of statistically significant features were identified. These features provide insight into metabolic behavior. For example, compounds that are abundant in the media but reduced in biological samples are likely being consumed by the organisms. Conversely, compounds that are elevated in specific cultures may represent metabolites produced by one organism or influenced by microbial interaction.
The reproducibility of the experimental setup is supported by consistent signal patterns across replicates, indicating reliable sample preparation and data acquisition.
Statistical Evaluation
To further refine the interpretation, pairwise statistical comparisons were performed between all sample classes. This enables identification of specific differences between monocultures and co-cultures, highlighting which metabolites are associated with individual organisms and which are affected by their interaction.
The results suggest that certain metabolites are dominant in Pseudomonas aeruginosa monocultures but are altered in co-culture conditions, indicating potential biological competition or suppression mechanisms.
Alternative Approach for Limited Data
Recognizing that biological replicates are not always available in practice, the study also explores an alternative strategy based on the coefficient of variation (CV). This method allows meaningful analysis even when only a single replicate per condition is available, making it particularly valuable for high-throughput or resource-limited experiments.
By applying this approach, the researchers were able to identify metabolites showing significant changes across conditions, including compounds uniquely produced or suppressed in specific cultures. One example includes the identification of 1-octen-3-ol, a known fungal metabolite, which serves as validation of the analytical workflow.
Discussion
A key outcome of this work is the demonstration that the choice of statistical method must be aligned with the experimental design. When sufficient replicates are available, variance-based methods provide robust results. In contrast, when data are limited, alternative metrics such as the coefficient of variation offer a practical solution. Additional approaches, such as fold-change analysis, can further enhance interpretation by focusing on relative differences between conditions.
The tile-based framework proves particularly effective for handling the complexity of GC×GC–MS data, enabling the extraction of meaningful biological information without compromising resolution.
Conclusion
This study highlights the power of advanced data processing strategies in modern analytical chemistry. Tile-based statistical analysis enables the systematic exploration of highly complex datasets and provides valuable insights into microbial metabolism and interaction. The approach is flexible, scalable, and adaptable to different experimental constraints, making it a strong candidate for broader application in metabolomics and environmental analysis.
Future Perspectives
Future work will focus on integrating chemometric tools to improve compound identification and expanding the analytical framework to support more comprehensive, multi-method data analysis.
This text has been automatically transcribed from a video presentation using AI technology. It may contain inaccuracies and is not guaranteed to be 100% correct.
-Workshop-LOGO_s.webp)
_s.webp)